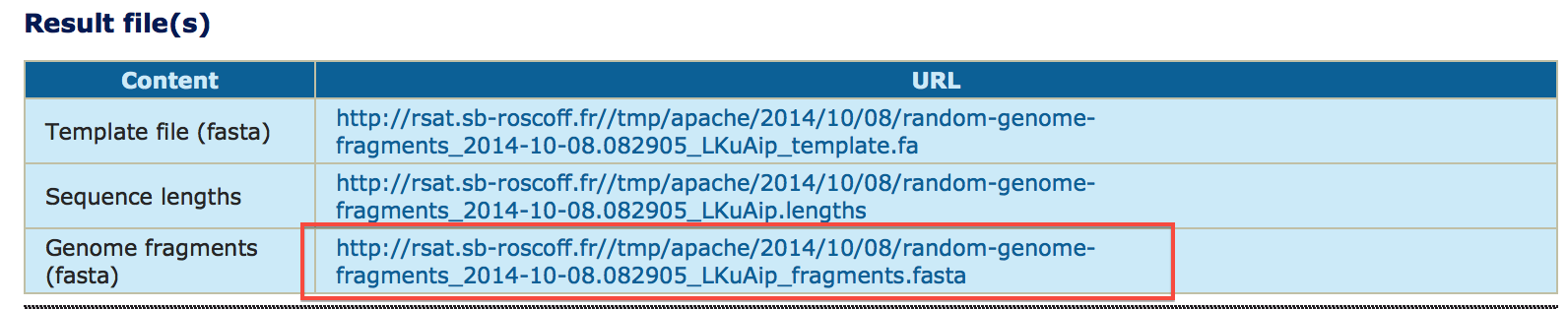
The program returns 3 links to files : the input BED file (coordinates), the corresponding sequences (FASTA file), and a log text file that contain information on the execution of the program.
Theodorou et al published a ChIP-seq experiment in the MCF7 cell line (breast cancer) to identify genomic locations bound by the transcription factor Estrogen receptor alpha (ESR1)(PMID:23172872). The particularity of this system is that it is inducible with the E2 (oestradiol) hormone. The authors were interested in evaluating the role of another transcription factor (GATA3) They thus performed the following experiments (with E2 induction):
| Peak BED file | Number of peaks | Sum of peak sizes (bp) |
|---|---|---|
| siGATA_ER_E2_r3_MACS_PeakSplitter.bed | 7.041 | 1.476.646 |
| siNT_ER_E2_r3_MACS_PeakSplitter.bed | 6.991 | 1.801.622 |
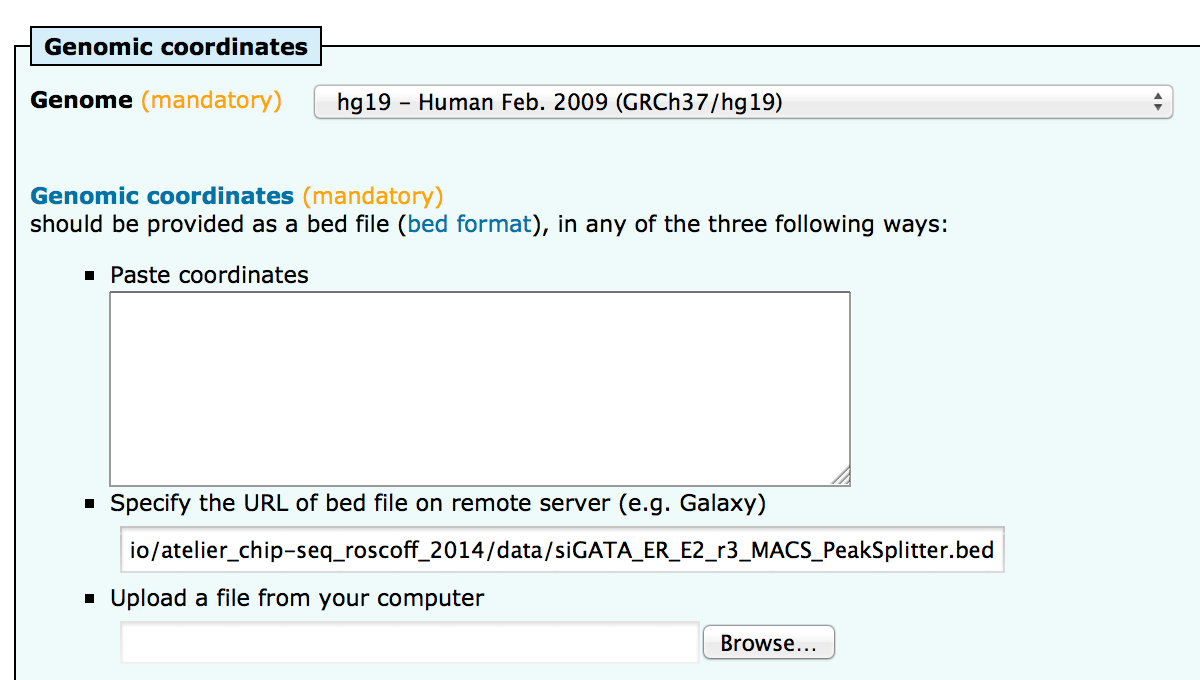

; sequences 7041All sequences were retrieved.
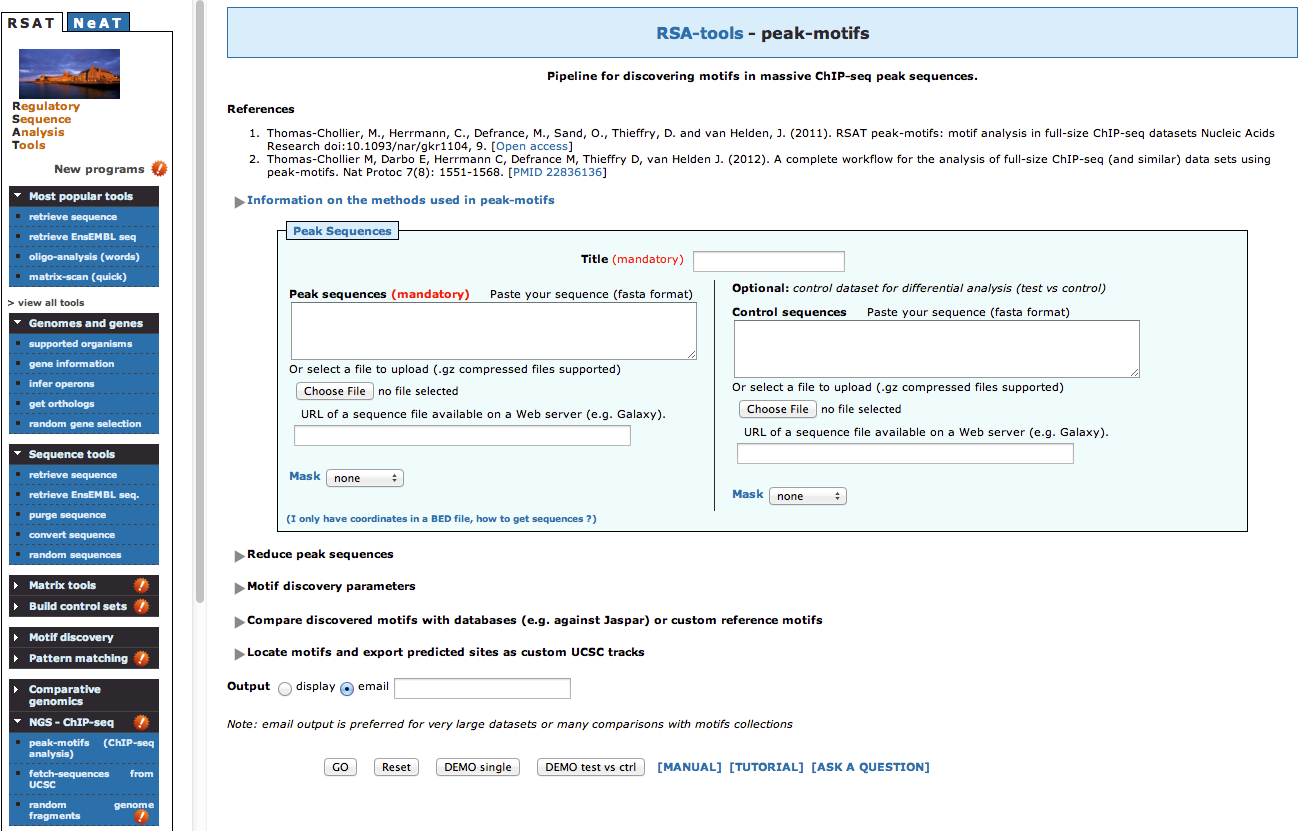
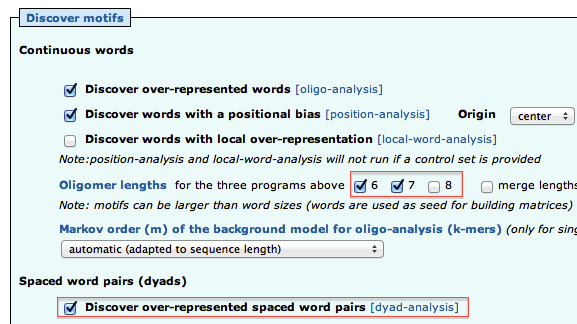

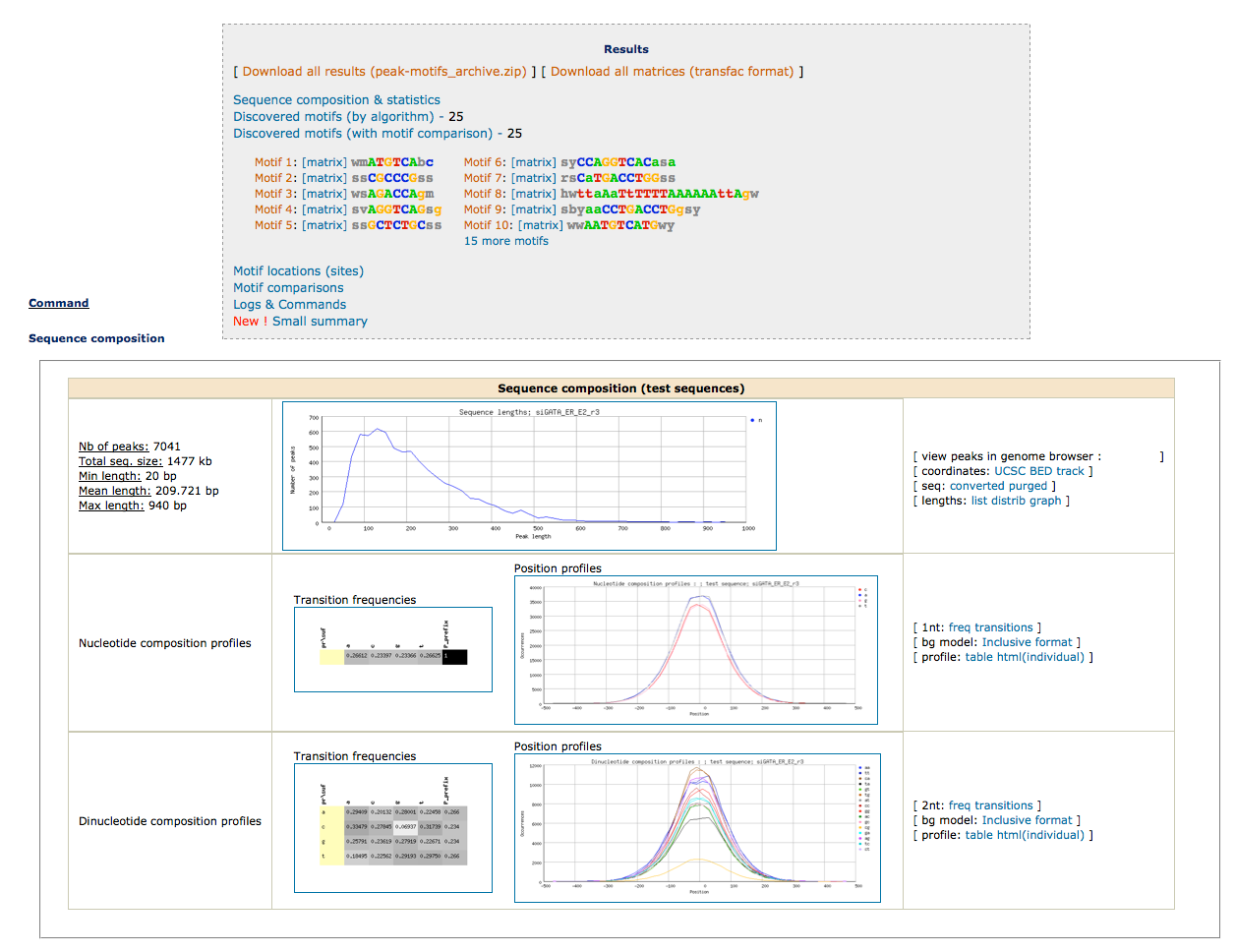
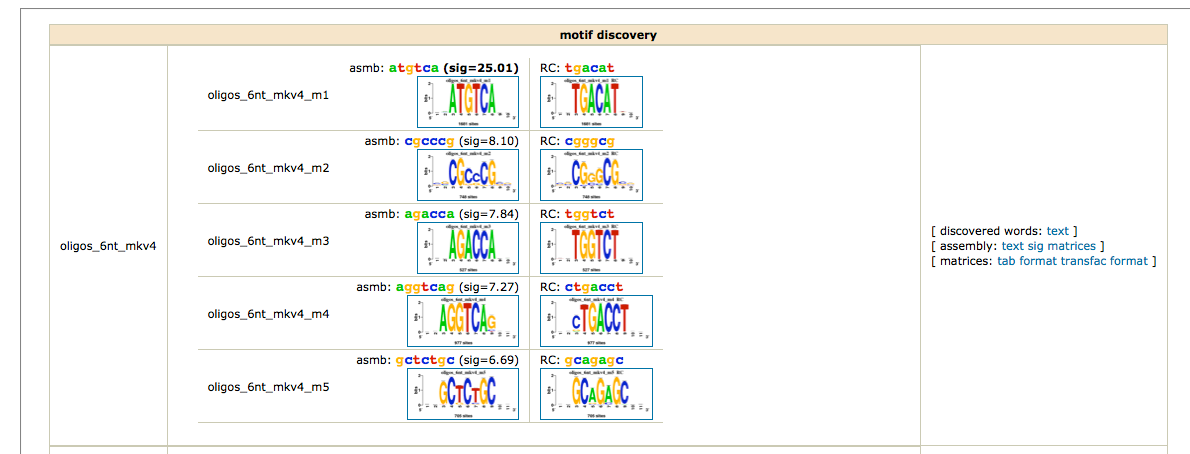
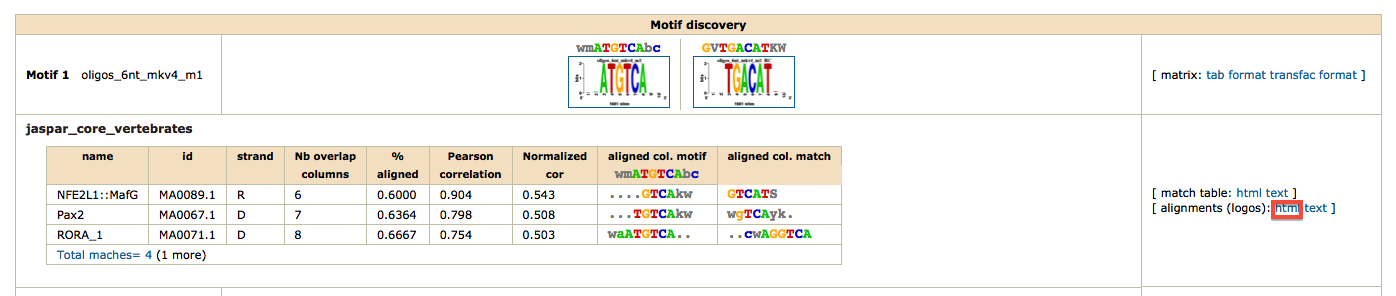
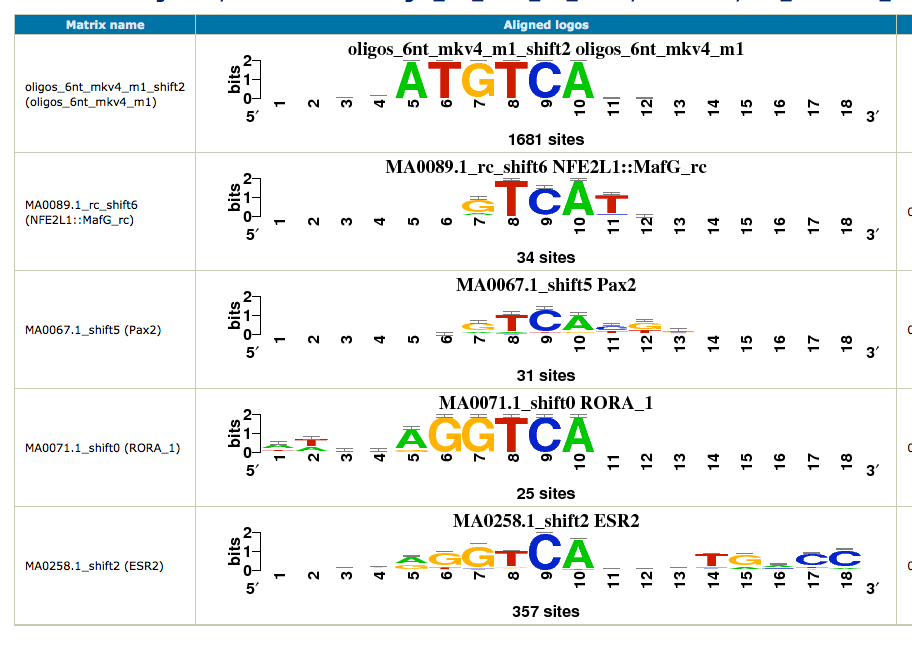
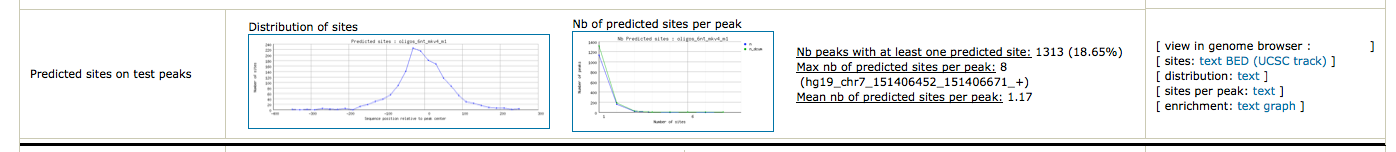
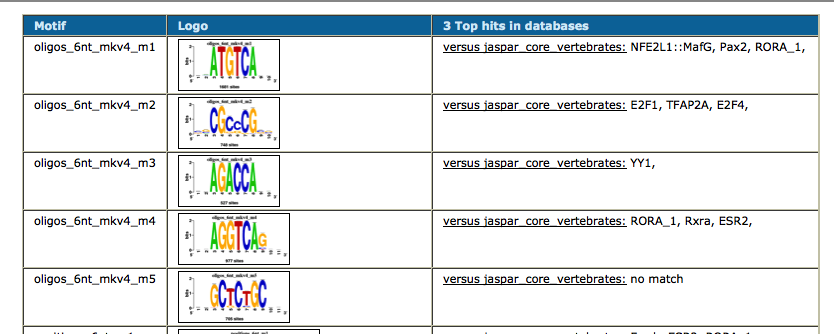
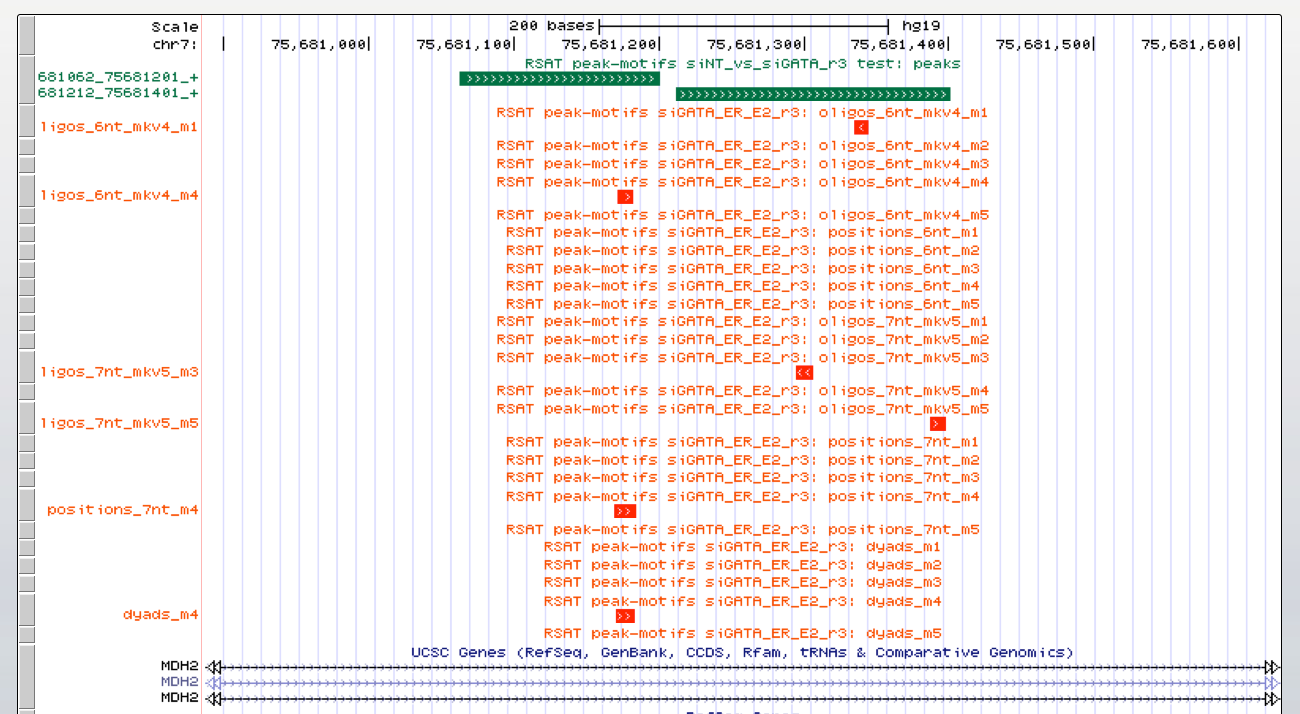
As we have two conditions (siGATA, siNT), we would like to find if some motifs are found in one dataset, but not the other. We are thus going to perform differential motif analysis.
As in experimental biology, we perform negative controls for motif analyses. RSAT has multiple options to create datasets that can serve as controls. Here, we will use "random genome fragments", to select random regions of the same number and size as the siGATA peak set.

Peak-motifs can prepare the BED files to be directly visualized in UCSC browser (or IGV !).
